csvtk - a cross-platform, efficient and practical CSV/TSV toolkit
- Documents: http://bioinf.shenwei.me/csvtk ( Usage, Tutorial and FAQs). 中文介绍
- Source code: https://github.com/shenwei356/csvtk
- Latest version:
Introduction
Similar to FASTA/Q format in field of Bioinformatics, CSV/TSV formats are basic and ubiquitous file formats in both Bioinformatics and data science.
People usually use spreadsheet software like MS Excel to process table data. However this is all by clicking and typing, which is not automated and is time-consuming to repeat, especially when you want to apply similar operations with different datasets or purposes.
You can also accomplish some CSV/TSV manipulations using shell commands, but more code is needed to handle the header line. Shell commands do not support selecting columns with column names either.
csvtk is convenient for rapid data investigation
and also easy to integrate into analysis pipelines.
It could save you lots of time in (not) writing Python/R scripts.
Table of Contents
- Introduction
- Table of Contents
- Features
- Subcommands
- Installation
- Command-line completion
- Compared to
csvkit - Examples
- Acknowledgements
- Contact
- License
- Starchart
Features
- Cross-platform (Linux/Windows/Mac OS X/OpenBSD/FreeBSD)
- Light weight and out-of-the-box, no dependencies, no compilation, no configuration
- Fast, multiple-CPUs supported (some commands)
- Practical functions provided by N subcommands
- Support STDIN and gzipped input/output file, easy being used in pipe
- Seamless support for xz (.xz), zstd (.zst), Bzip2 (.bz2), LZ4 (.lz4) formats
- Most of the subcommands support unselecting fields and fuzzy fields,
e.g.
-f "-id,-name"for all fields except "id" and "name",-F -f "a.*"for all fields with prefix "a.". - Support some common plots (see usage)
Seamless support for data with meta line (e.g.,sep=,) of separator declaration used by MS Excel
Subcommands
57 subcommands in total.
Information
headers: prints headersdim: dimensions of CSV filenrow: print number of recordsncol: print number of columnssummary: summary statistics of selected numeric or text fields (groupby group fields)watch: online monitoring and histogram of selected fieldcorr: calculate Pearson correlation between numeric columns
Format conversion
pretty: converts CSV to a readable aligned tablecsv2tab: converts CSV to tabular formattab2csv: converts tabular format to CSVspace2tab: converts space delimited format to TSVcsv2md: converts CSV to markdown formatcsv2rst: converts CSV to reStructuredText formatcsv2json: converts CSV to JSON formatcsv2xlsx: converts CSV/TSV files to XLSX filexlsx2csv: converts XLSX to CSV format
Set operations
head: prints first N recordsconcat: concatenates CSV/TSV files by rowssample: sampling by proportioncut: select and arrange fieldsgrep: greps data by selected fields with patterns/regular expressionsuniq: unique data without sortingfreq: frequencies of selected fieldsinter: intersection of multiple filesfilter: filters rows by values of selected fields with arithmetic expressionfilter2: filters rows by awk-like arithmetic/string expressionsjoin: join files by selected fields (inner, left and outer join)splitsplits CSV/TSV into multiple files according to column valuessplitxlsx: splits XLSX sheet into multiple sheets according to column valuescomb: compute combinations of items at every row
Edit
fix: fix CSV/TSV with different numbers of columns in rowsfix-quotes: fix malformed CSV/TSV caused by double-quotesdel-quotes: remove extra double-quotes added byfix-quotesadd-header: add column namesdel-header: delete column namesrename: renames column names with new namesrename2: renames column names by regular expressionreplace: replaces data of selected fields by regular expressionround: round float to n decimal placescomma: make numbers more readable by adding commasmutate: creates new columns from selected fields by regular expressionmutate2: creates a new column from selected fields by awk-like arithmetic/string expressionsmutate3: create a new column from selected fields with Go-like expressionsfmtdate: format date of selected fields
Transform
transpose: transposes CSV datasep: separate column into multiple columnsgather: gather columns into key-value pairs, liketidyr::gather/pivot_longerspread: spread a key-value pair across multiple columns, liketidyr::spread/pivot_widerunfold: unfold multiple values in cells of a fieldfold: fold multiple values of a field into cells of groups
Ordering
Ploting
Misc
catstream file and report progressversionprint version information and check for updategenautocompletegenerate shell autocompletion script (bash|zsh|fish|powershell)
Installation
csvtk is implemented in Go programming language,
executable binary files for most popular operating systems are freely available
in release page.
Method 1: Download binaries (latest stable/dev version)
Just download compressed
executable file of your operating system,
and decompress it with tar -zxvf *.tar.gz command or other tools.
And then:
-
For Linux-like systems
-
If you have root privilege simply copy it to
/usr/local/bin:sudo cp csvtk /usr/local/bin/ -
Or copy to anywhere in the environment variable
PATH:mkdir -p $HOME/bin/; cp csvtk $HOME/bin/
-
-
For windows, just copy
csvtk.exetoC:\WINDOWS\system32.
Method 2: Install via Pixi
pixi global install csvtk
Method 3: Install via conda (latest stable version) 
# >= v0.31.0
conda install -c conda-forge csvtk
# <= v0.31.0
conda install -c bioconda csvtk
Method 4: Install via homebrew
brew install csvtk
Method 5: For Go developer (latest stable/dev version)
go install github.com/shenwei356/csvtk/csvtk@latest
Method 6: For ArchLinux AUR users (may be not the latest)
yaourt -S csvtk
Command-line completion
Bash:
# generate completion shell
csvtk genautocomplete --shell bash
# configure if never did.
# install bash-completion if the "complete" command is not found.
echo "for bcfile in ~/.bash_completion.d/* ; do source \$bcfile; done" >> ~/.bash_completion
echo "source ~/.bash_completion" >> ~/.bashrc
Zsh:
# generate completion shell
csvtk genautocomplete --shell zsh --file ~/.zfunc/_csvtk
# configure if never did
echo 'fpath=( ~/.zfunc "${fpath[@]}" )' >> ~/.zshrc
echo "autoload -U compinit; compinit" >> ~/.zshrc
fish:
csvtk genautocomplete --shell fish --file ~/.config/fish/completions/csvtk.fish
Compared to csvkit
csvkit, attention: this table wasn't updated for many years.
| Features | csvtk | csvkit | Note |
|---|---|---|---|
| Read Gzip | Yes | Yes | read gzip files |
| Fields ranges | Yes | Yes | e.g. -f 1-4,6 |
| Unselect fields | Yes | -- | e.g. -1 for excluding first column |
| Fuzzy fields | Yes | -- | e.g. ab* for columns with name prefix "ab" |
| Reorder fields | Yes | Yes | it means -f 1,2 is different from -f 2,1 |
| Rename columns | Yes | -- | rename with new name(s) or from existed names |
| Sort by multiple keys | Yes | Yes | bash sort like operations |
| Sort by number | Yes | -- | e.g. -k 1:n |
| Multiple sort | Yes | -- | e.g. -k 2:r -k 1:nr |
| Pretty output | Yes | Yes | convert CSV to readable aligned table |
| Unique data | Yes | -- | unique data of selected fields |
| frequency | Yes | -- | frequencies of selected fields |
| Sampling | Yes | -- | sampling by proportion |
| Mutate fields | Yes | -- | create new columns from selected fields |
| Replace | Yes | -- | replace data of selected fields |
Similar tools:
- csvkit - A suite of utilities for converting to and working with CSV, the king of tabular file formats. http://csvkit.rtfd.org/
- xsv - A fast CSV toolkit written in Rust.
- miller - Miller is like sed, awk, cut, join, and sort for name-indexed data such as CSV and tabular JSON http://johnkerl.org/miller
- tsv-utils - Command line utilities for tab-separated value files written in the D programming language.
Examples
Attention
- By default, csvtk assumes input files have header row, if not, switch flag
-Hon. - By default, csvtk handles CSV files, use flag
-tfor tab-delimited files. - Column names should be unique.
- By default, lines starting with
#will be ignored, if the header row starts with#, please assign flag-Canother rare symbol, e.g.$. - Do not mix use field (column) numbers and names to specify columns to operate.
- The CSV parser requires all the lines have same numbers of fields/columns.
Even lines with spaces will cause error.
Use
-I/--ignore-illegal-rowto skip these lines if neccessary. You can also use "csvtk fix" to fix files with different numbers of columns in rows. -
If double-quotes exist in fields not enclosed with double-quotes, e.g.,
x,a "b" c,1It would report error:
bare `"` in non-quoted-field.Please switch on the flag
-lor usecsvtk fix-quotesto fix it. -
If somes fields have only a double-quote either in the beginning or in the end, e.g.,
x,d "e","a" b c,1It would report an error:
extraneous or missing " in quoted-fieldPlease use
csvtk fix-quotesto fix it, and usecsvtk del-quotesto reset to the original format as needed.
Examples
-
Pretty result
$ csvtk pretty names.csv id first_name last_name username -- ---------- --------- -------- 11 Rob Pike rob 2 Ken Thompson ken 4 Robert Griesemer gri 1 Robert Thompson abc NA Robert Abel 123 $ csvtk pretty names.csv -S 3line ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ id first_name last_name username ──────────────────────────────────────── 11 Rob Pike rob 2 Ken Thompson ken 4 Robert Griesemer gri 1 Robert Thompson abc NA Robert Abel 123 ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ $ csvtk pretty names.csv -S round -w 5 -m 1- ╭───────┬────────────┬───────────┬──────────╮ │ id │ first_name │ last_name │ username │ ├───────┼────────────┼───────────┼──────────┤ │ 11 │ Rob │ Pike │ rob │ ├───────┼────────────┼───────────┼──────────┤ │ 2 │ Ken │ Thompson │ ken │ ├───────┼────────────┼───────────┼──────────┤ │ 4 │ Robert │ Griesemer │ gri │ ├───────┼────────────┼───────────┼──────────┤ │ 1 │ Robert │ Thompson │ abc │ ├───────┼────────────┼───────────┼──────────┤ │ NA │ Robert │ Abel │ 123 │ ╰───────┴────────────┴───────────┴──────────╯ -
Summary of selected numeric fields, supporting "group-by"
$ cat testdata/digitals2.csv \ | csvtk summary -i -f f4:sum,f5:sum -g f1,f2 \ | csvtk pretty f1 f2 f4:sum f5:sum bar xyz 7.00 106.00 bar xyz2 4.00 4.00 foo bar 6.00 3.00 foo bar2 4.50 5.00 -
Select fields/columns (
cut)- By index:
csvtk cut -f 1,2 - By names:
csvtk cut -f first_name,username - Unselect:
csvtk cut -f -1,-2orcsvtk cut -f -first_name - Fuzzy fields:
csvtk cut -F -f "*_name,username" - Field ranges:
csvtk cut -f 2-4for column 2,3,4 orcsvtk cut -f -3--1for discarding column 1,2,3 - All fields:
csvtk cut -f 1-orcsvtk cut -F -f "*"
- By index:
-
Search by selected fields (
grep) (matched parts will be highlighted as red)- By exactly matching:
csvtk grep -f first_name -p Robert -p Rob - By regular expression:
csvtk grep -f first_name -r -p Rob - By pattern list:
csvtk grep -f first_name -P name_list.txt - Remore rows containing missing data (NA):
csvtk grep -F -f "*" -r -p "^$" -v
- By exactly matching:
-
Rename column names (
renameandrename2)- Setting new names:
csvtk rename -f A,B -n a,borcsvtk rename -f 1-3 -n a,b,c - Replacing with original names by regular express:
csvtk rename2 -f 1- -p "(.*)" -r 'prefix_$1'for adding prefix to all column names.
- Setting new names:
-
Edit data with regular expression (
replace)- Remove Chinese charactors:
csvtk replace -F -f "*_name" -p "\p{Han}+" -r ""
- Remove Chinese charactors:
-
Create new column from selected fields by regular expression (
mutate)- In default, copy a column:
csvtk mutate -f id - Extract prefix of data as group name (get "A" from "A.1" as group name):
csvtk mutate -f sample -n group -p "^(.+?)\." --after sample
- In default, copy a column:
-
Sort by multiple keys (
sort)- By single column :
csvtk sort -k 1orcsvtk sort -k last_name - By multiple columns:
csvtk sort -k 1,2orcsvtk sort -k 1 -k 2orcsvtk sort -k last_name,age - Sort by number:
csvtk sort -k 1:norcsvtk sort -k 1:nrfor reverse number - Complex sort:
csvtk sort -k region -k age:n -k id:nr - In natural order:
csvtk sort -k chr:N
- By single column :
-
Join multiple files by keys (
join)- All files have same key column:
csvtk join -f id file1.csv file2.csv - Files have different key columns:
csvtk join -f "username;username;name" names.csv phone.csv adress.csv -k
- All files have same key column:
-
Filter by numbers (
filter)- Single field:
csvtk filter -f "id>0" - Multiple fields:
csvtk filter -f "1-3>0" - Using
--anyto print record if any of the field satisfy the condition:csvtk filter -f "1-3>0" --any - fuzzy fields:
csvtk filter -F -f "A*!=0"
- Single field:
-
Filter rows by awk-like arithmetic/string expressions (
filter2)- Using field index:
csvtk filter2 -f '$3>0' - Using column names:
csvtk filter2 -f '$id > 0' - Both arithmetic and string expressions:
csvtk filter2 -f '$id > 3 || $username=="ken"' - More complicated:
csvtk filter2 -H -t -f '$1 > 2 && $2 % 2 == 0'
- Using field index:
-
Plotting
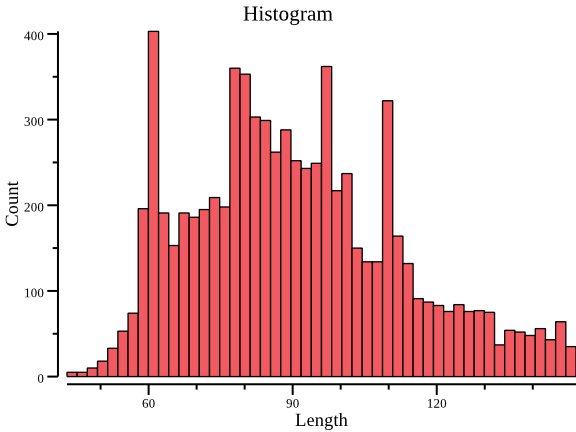
- plot histogram with data of the second column:
csvtk -t plot hist testdata/grouped_data.tsv.gz -f 2 | display
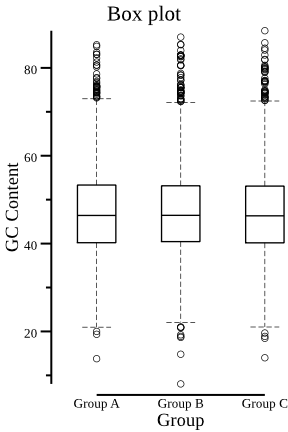
- plot boxplot with data of the "GC Content" (third) column,
group information is the "Group" column.
csvtk -t plot box testdata/grouped_data.tsv.gz -g "Group" \ -f "GC Content" --width 3 --title "Box plot" | display
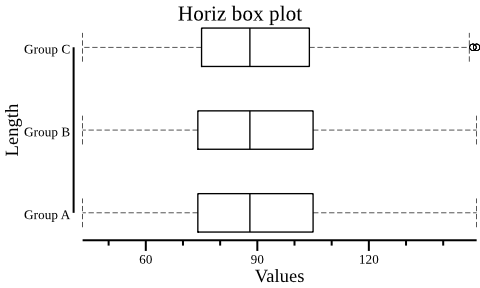
- plot horiz boxplot with data of the "Length" (second) column,
group information is the "Group" column.
csvtk -t plot box testdata/grouped_data.tsv.gz -g "Group" -f "Length" \ --height 3 --width 5 --horiz --title "Horiz box plot" | display
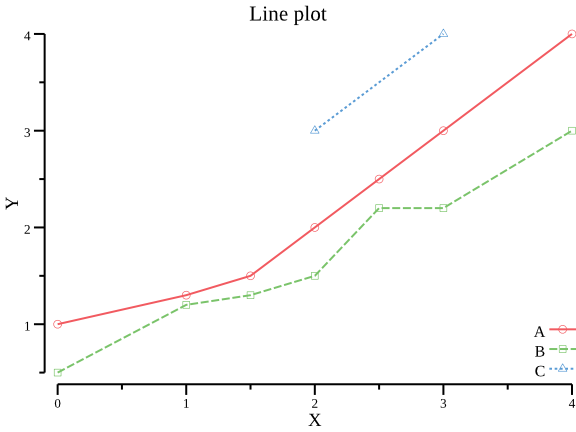
- plot line plot with X-Y data
csvtk -t plot line testdata/xy.tsv -x X -y Y -g Group | display
- plot line plot with X-Y data (where X values are dates or other sortable values)
cat testdata/date2value.csv \ | csvtk gather -f 2- -k type -v count \ | csvtk plot line --group-field type -x date -y count --data-field-x-nominal \ -o testdata/figures/line_plot_date.png

- plot scatter plot with X-Y data
csvtk -t plot line testdata/xy.tsv -x X -y Y -g Group --scatter | display
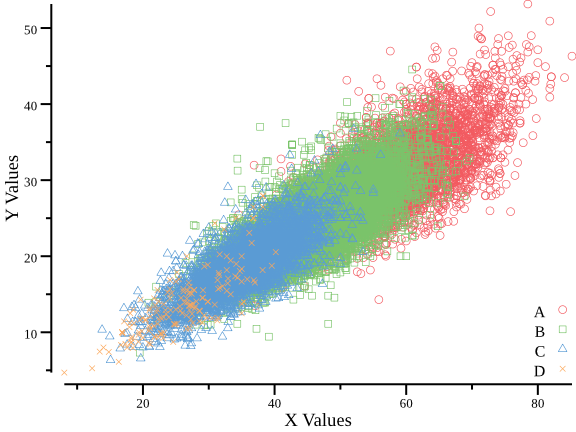
- plot histogram with data of the second column:
Acknowledgements
We are grateful to Zhiluo Deng and Li Peng for suggesting features and reporting bugs.
Thanks Albert Vilella for feature suggestions, which makes csvtk feature-rich。
Contact
Create an issue to report bugs, propose new functions or ask for help.
Or leave a comment.




